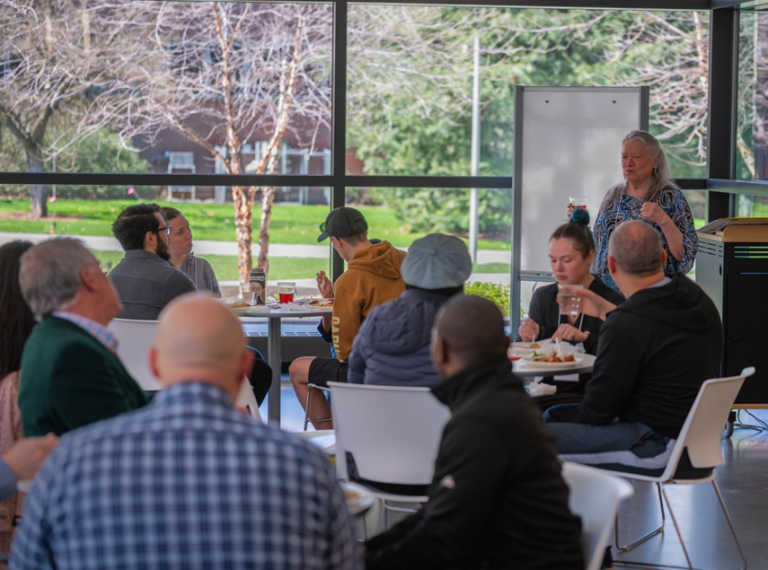
Bridging Disciplines and Building Teams: A Day of Collaboration and Research
Written by Chali MintoPhotos by Travis Warner In a vibrant display of cross-campus collaboration, colleagues from all over the University of Idaho campus joined us for a unique event focused on fostering new connections and generating innovative and collaborative research ideas. Faculty and staff from diverse disciplines—including biology, engineering, and law—gathered to build teams with…